Explained: Singular Value Decomposition (SVD)
In which I attempt an intuitive explanation of singular value decomposition (SVD).

SVD is a matrix decomposition method for decomposition a matrix into smaller parts. According to SVD, a matrix can be broken down and written as the dot product of three additional matrices, which have some nice properties. For example, one of these matrices is a condensed representation of the original matrix that is generally useful in quite a few contexts. More on this later. These matrices are given by the formula below, where A is the dataset being decomposed with SVD, and contains x samples and d features.
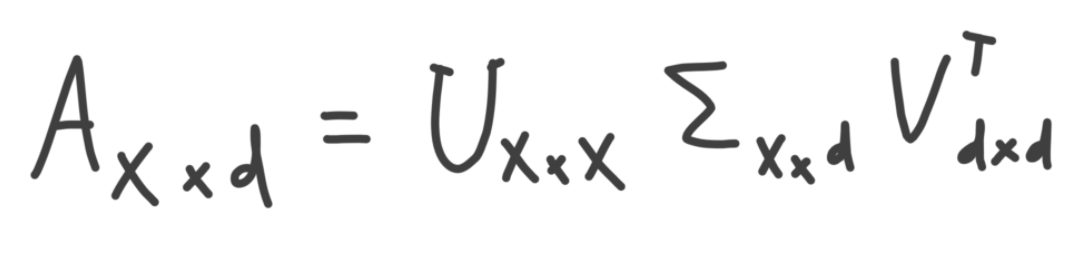
I'll swing back around to this formula, but first I'd like to build an intuition for what some of the above matrices are.
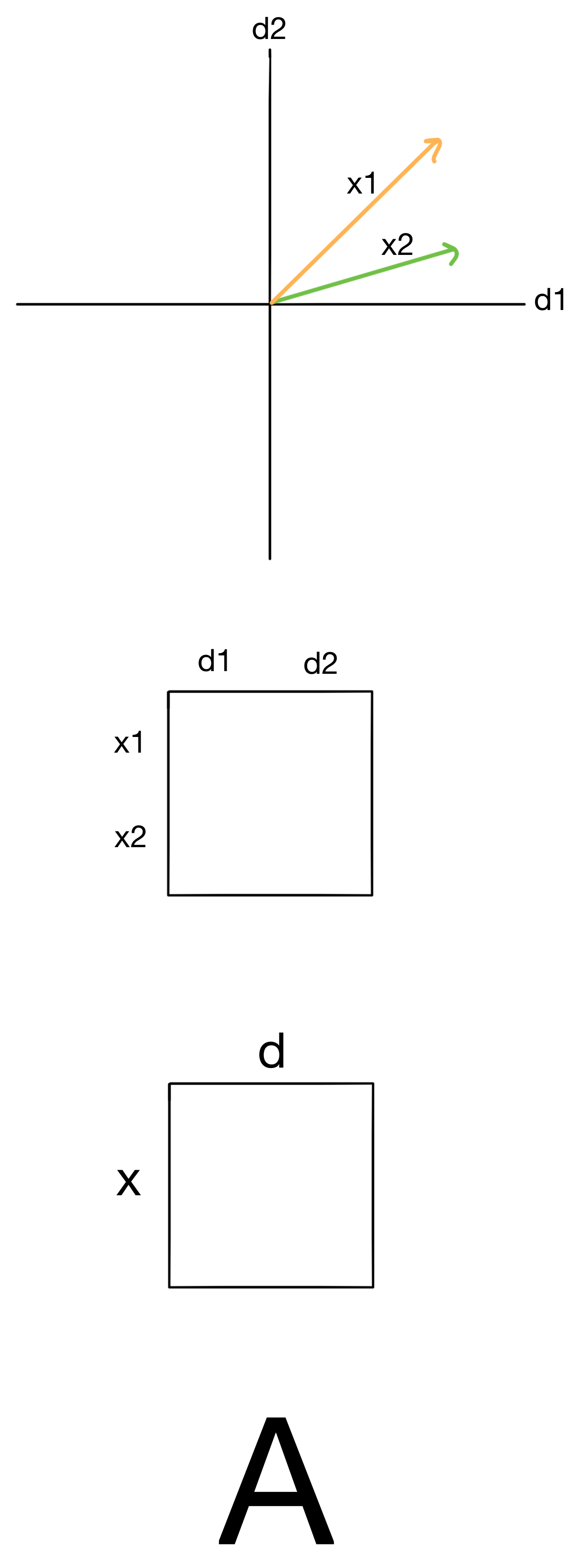
I'll be working through some of the below examples with two datasets: one with two samples, x1 and x2, and two features, d1 and d2 (for visualization purposes), and another which has x samples and d features. I call both these datasets A. I'll be walking through these in parallel to hopefully build an intuition for whats going on in both low and high dimensional space. We have vectors plotted for the smaller dataset (top), the matrix representing the smaller dataset (middle), and the matrix representing our larger dataset (bottom).
Courtesy of some matrix operations, we can derive the nifty formula shown below.
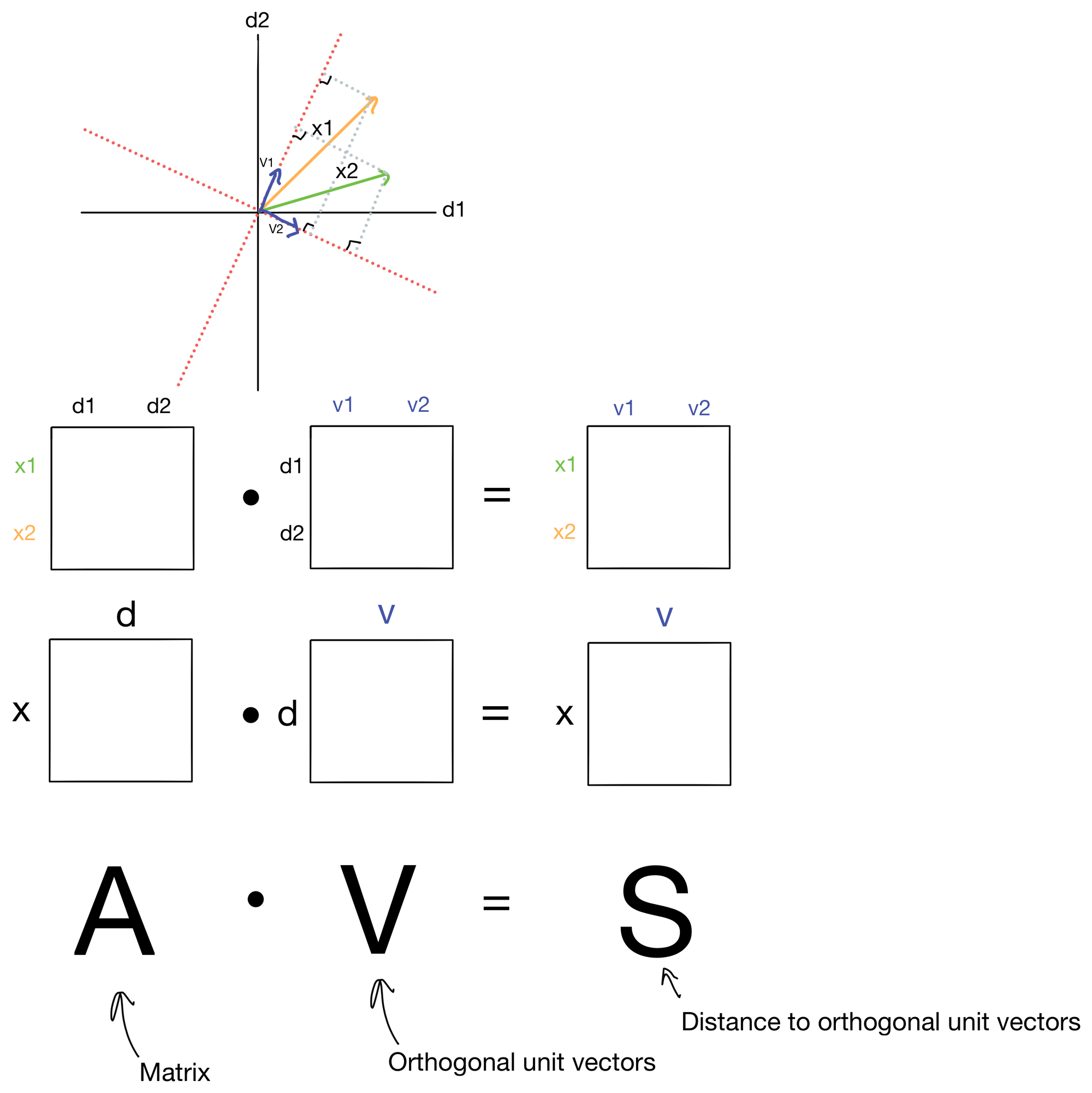
First, we define a set (in blue) of orthonormal basis vectors v1 and v2. For now don't worry about how these orthonormal vectors are generated, just know that they are orthogonal (i.e. 90 degrees from each other) and normal (have a magnitude of 1). The number of basis vectors will equal the number of features in the dataset, i.e. d=v. We can then use A to transform these basis vectors by taking the dot product of the two matrices. The entries in the resulting matrix S contain the distances from a given sample vector in A to each basis vector v. These distances are shown in gray on the plot.
With some algebra, this equation can actually be rewritten.
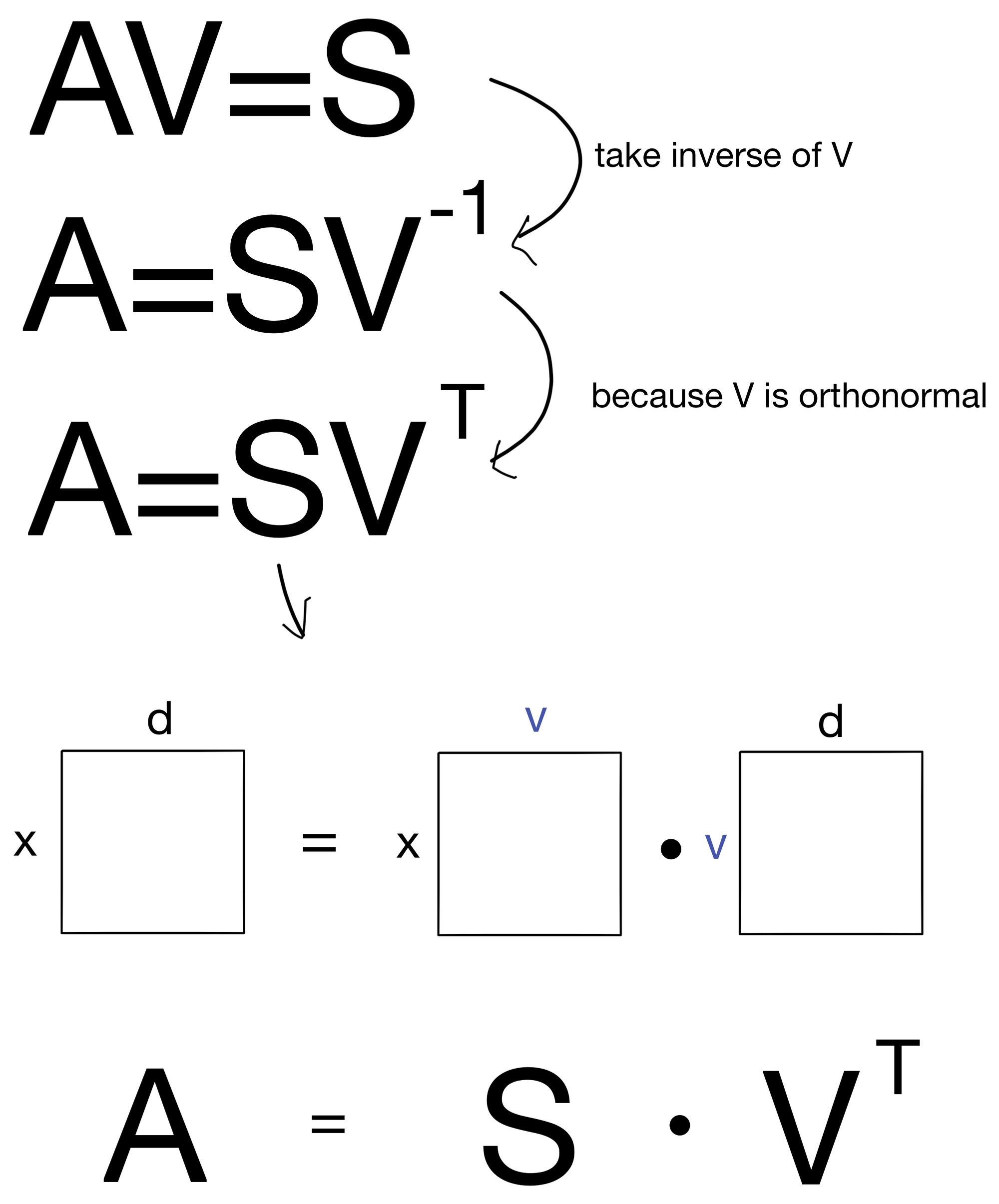
Now we have A on one side and S and VT on the other. As we see below, this is half way to our target formula!
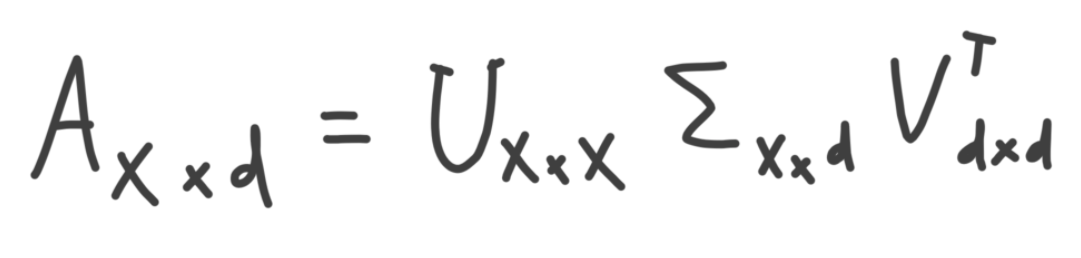
All we need to do is split S into two new terms.
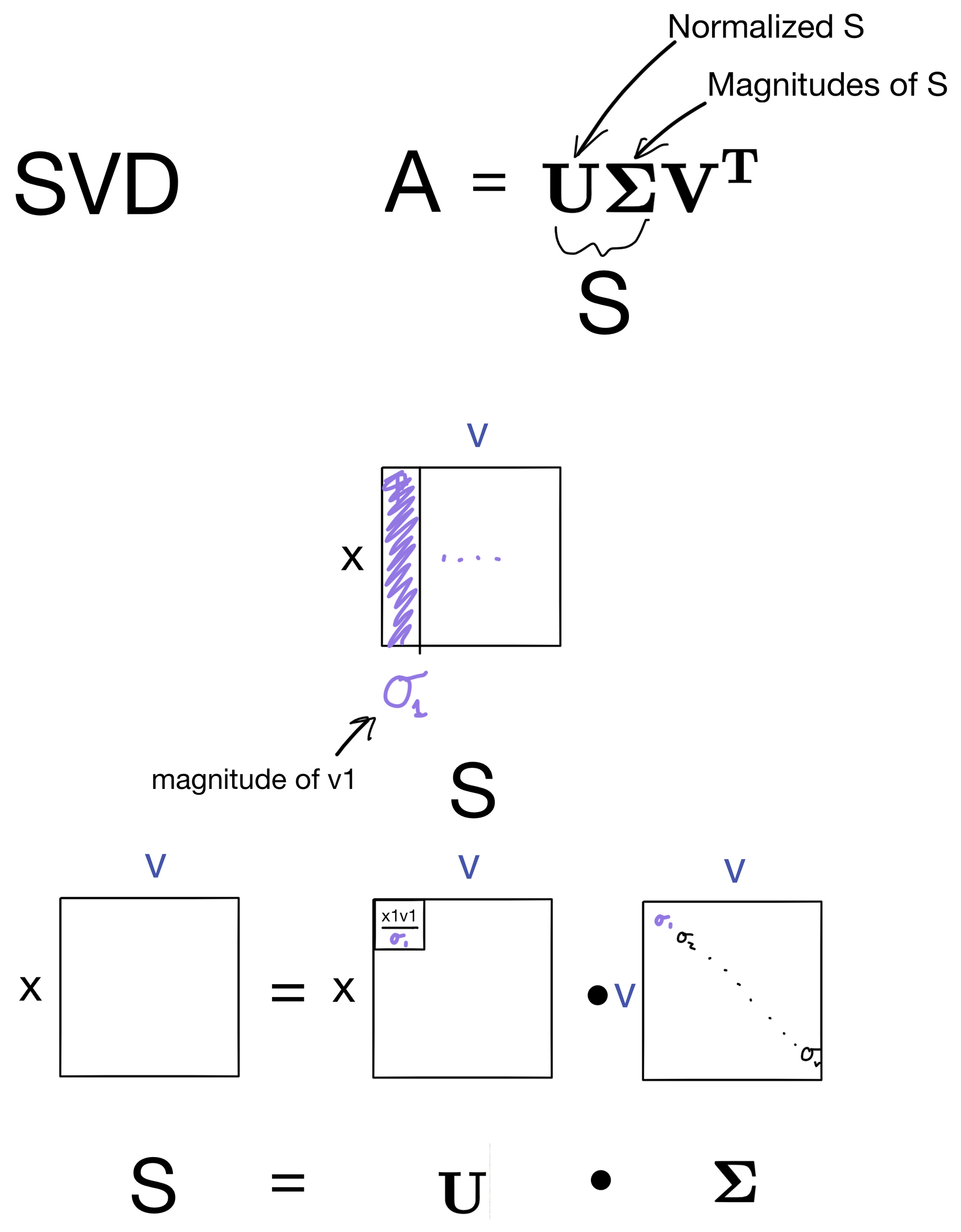
When we split S, we do it in such a way that the matrix U is a normalized version of S, where each vector in S is a unit vector with magnitude of 1. In other words, U contains the direction of S, but not the magnitude. Meaning that to reconstruct S, E will need to be a diagonal matrix where each entry on the diagonal are magnitudes of the directional vectors in U. This deconstruction is shown above.
Nearly there. We can now write the (nearly, as the dimensions don't quite fully match up yet) full version of the formula.
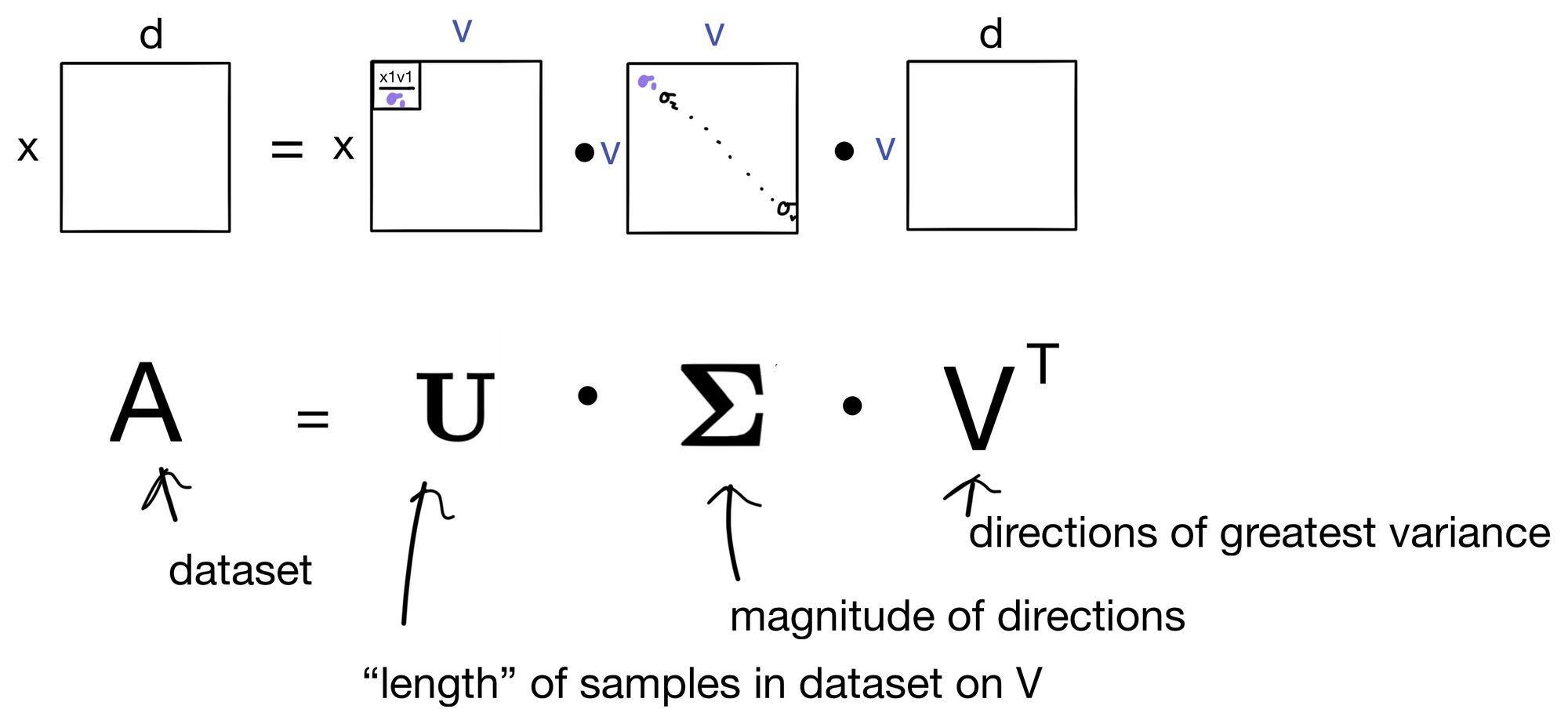
Additionally, if we look back we can label the matrices with their thematic representations, which is why SVD is so useful. U is an "information rich" version of A, where if we take the first n columns they will contain a condensed representation of the dataset. The magnitudes in E are rank ordered and correspond to the columns in U, and can be thought of as the importance for each column in U. Finally, VT contains the basis vectors which give the direction of greatest variance in the dataset. Another description would be the directions that contain the most information about the dataset A. This is ultimately why U is an information rich version of A. For more about this concept, and how the basis vectors are derived check out my post on principle component analysis (the principle components are VT in this context, although it's important that they are derived differently than they are in SVD - they just end up being the same thing).
But we aren't quite finished yet, the dimensionality of U and E is a little off in comparison the the SVD formulation below.
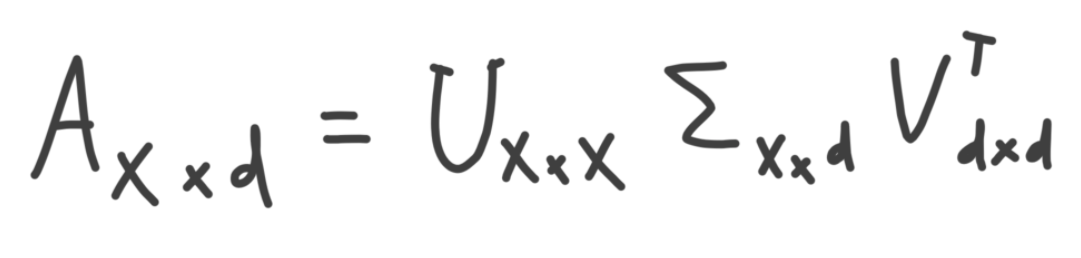
This is because in SVD the diagonal matrix E will contain zero padding. Either on the bottom if x > d (shown below), or the side if x < d. So when you take the dot product UE you ultimately end up with the relevant information confined to a size d x d matrix. Also, remember from above that v = d.
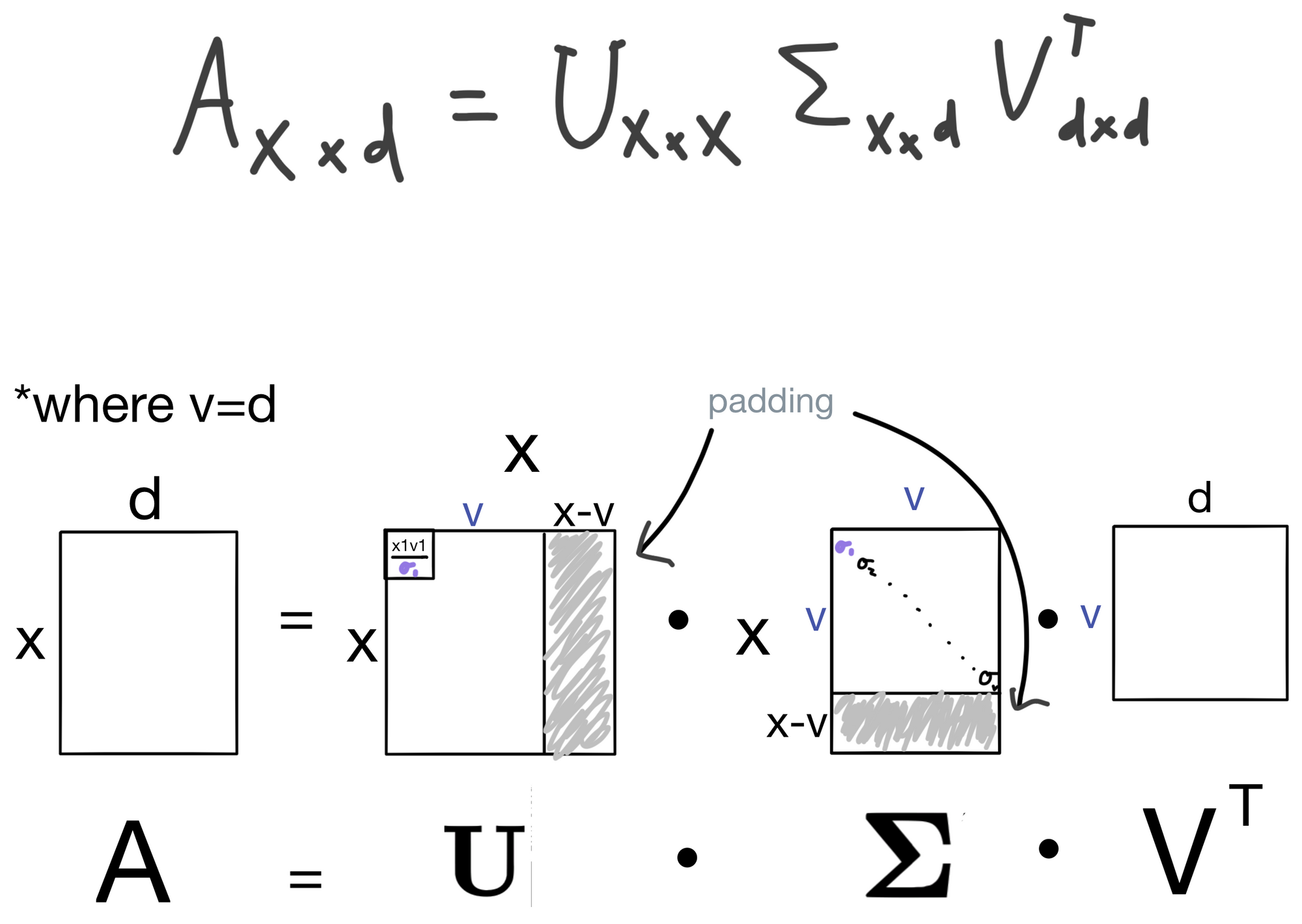
There you have it, a (hopefully) intuitive explanation for what SVD is and why it's useful.
H/T to Gregory Gunderson and Reza Bagheri where inspiration for portions of this post was taken.
Cheers :)
